3D Nuclei Clustering Tool
- Details
- MRI tools
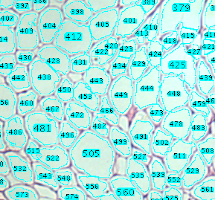
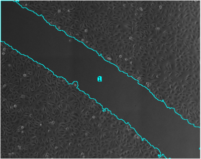
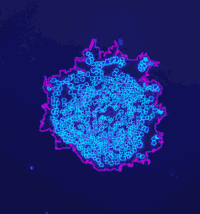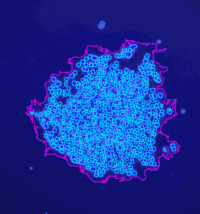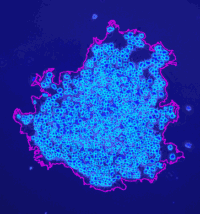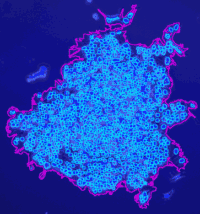
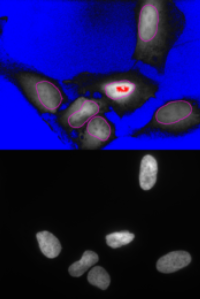
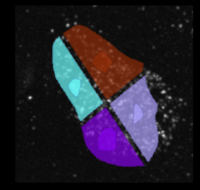
Analyze the clustering behavior of nuclei in 3D images. The centers of the nuclei are detected. The nuclei are filtered by the presence of a signal in a different channel. They clustered with the density based algorithm DBSCAN. The nearest neighbor distances between all nuclei and those outside and inside of the clusters are calculated.
Download and further informations: 3D_Nuclei_Clustering_Tool on github